This solution accelerator is aimed at Computational Biologists working with genomic data. It aims to integrate genomics data with other relevant datasets, presenting findings through interactive dashboards for clinical scientists, geneticists, and other practitioners. Our focus lies in analyzing population-level trends and identifying samples associated with specific causal variants previously discovered through Genome Wide Association Studies (GWAS). We utilize the GWAS catalog in this accelerator.
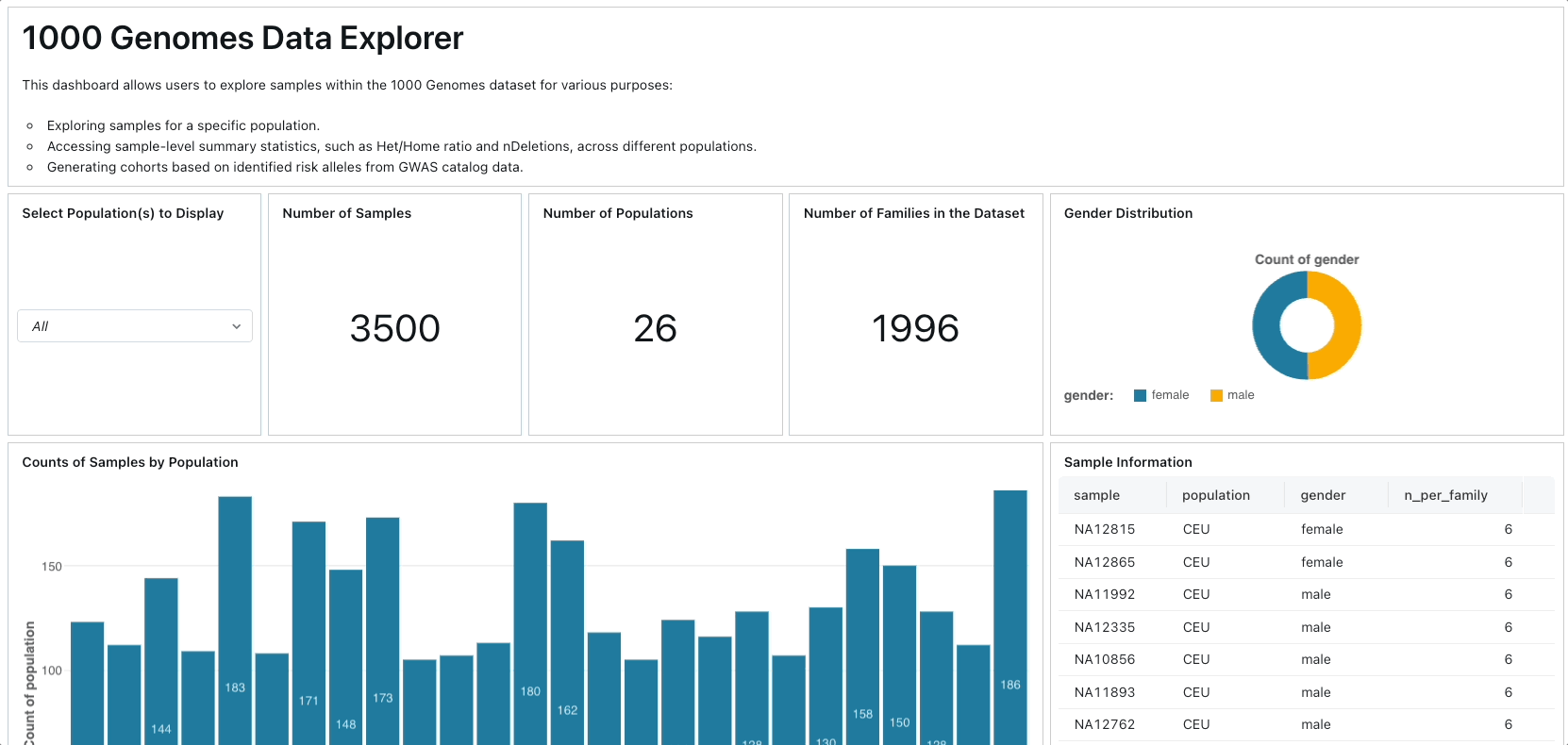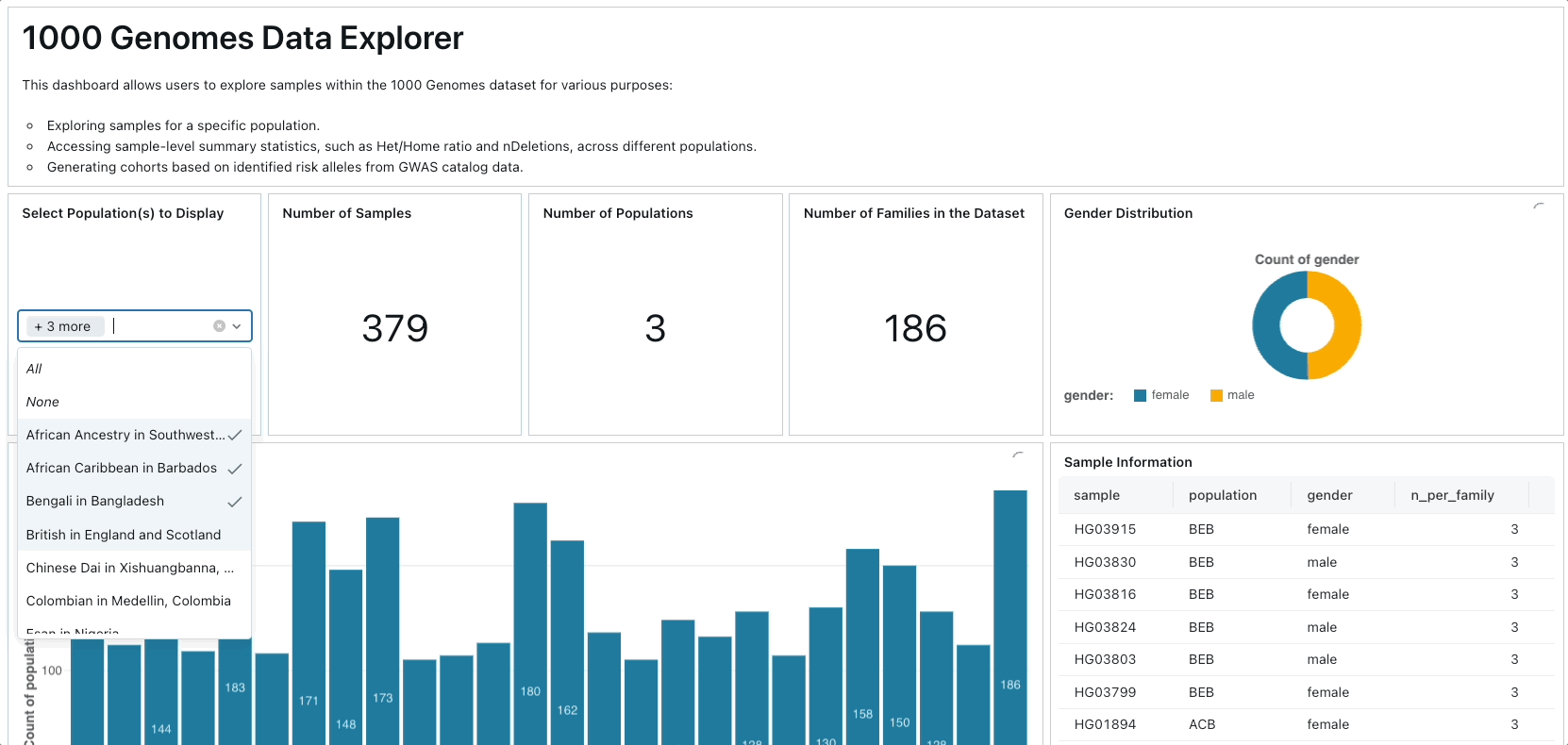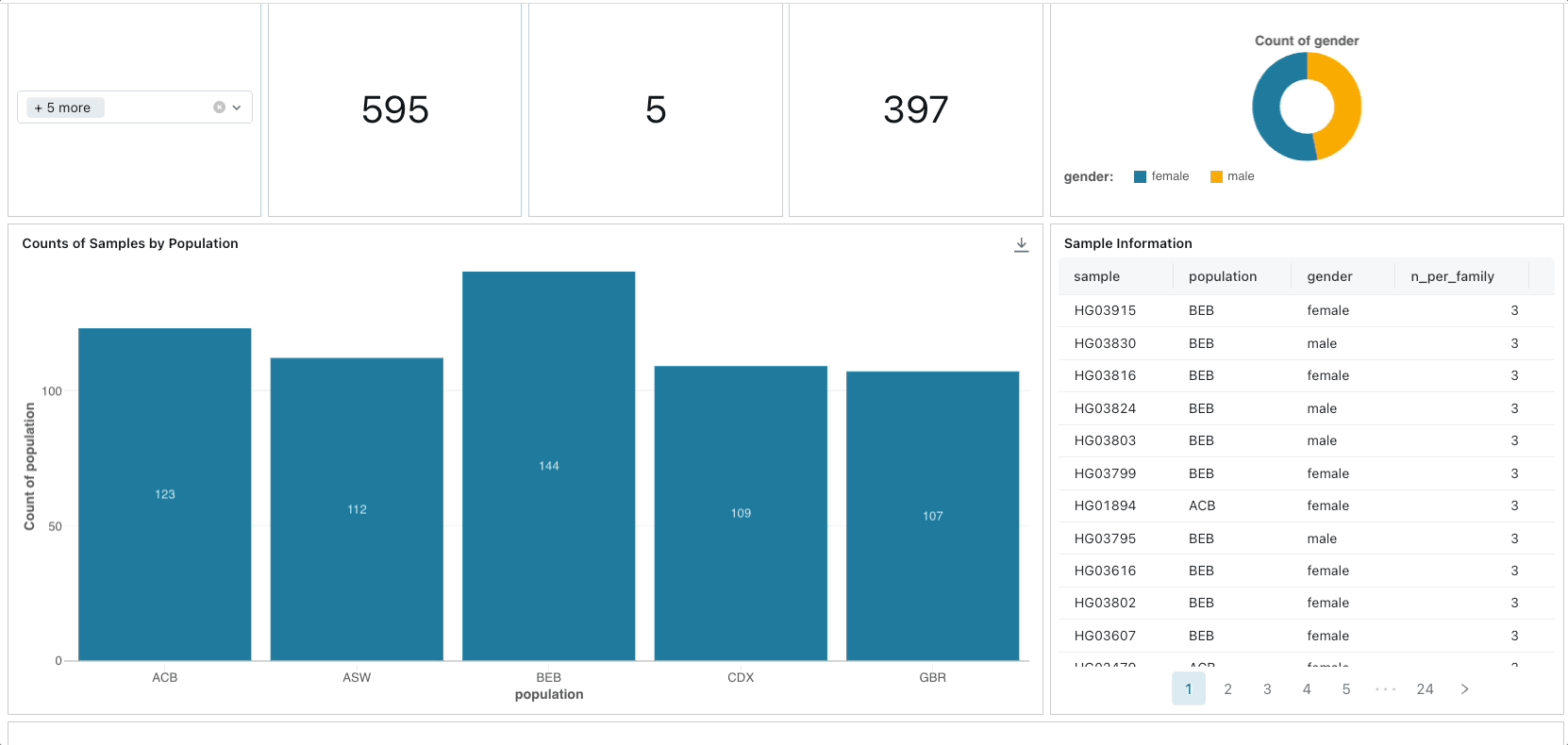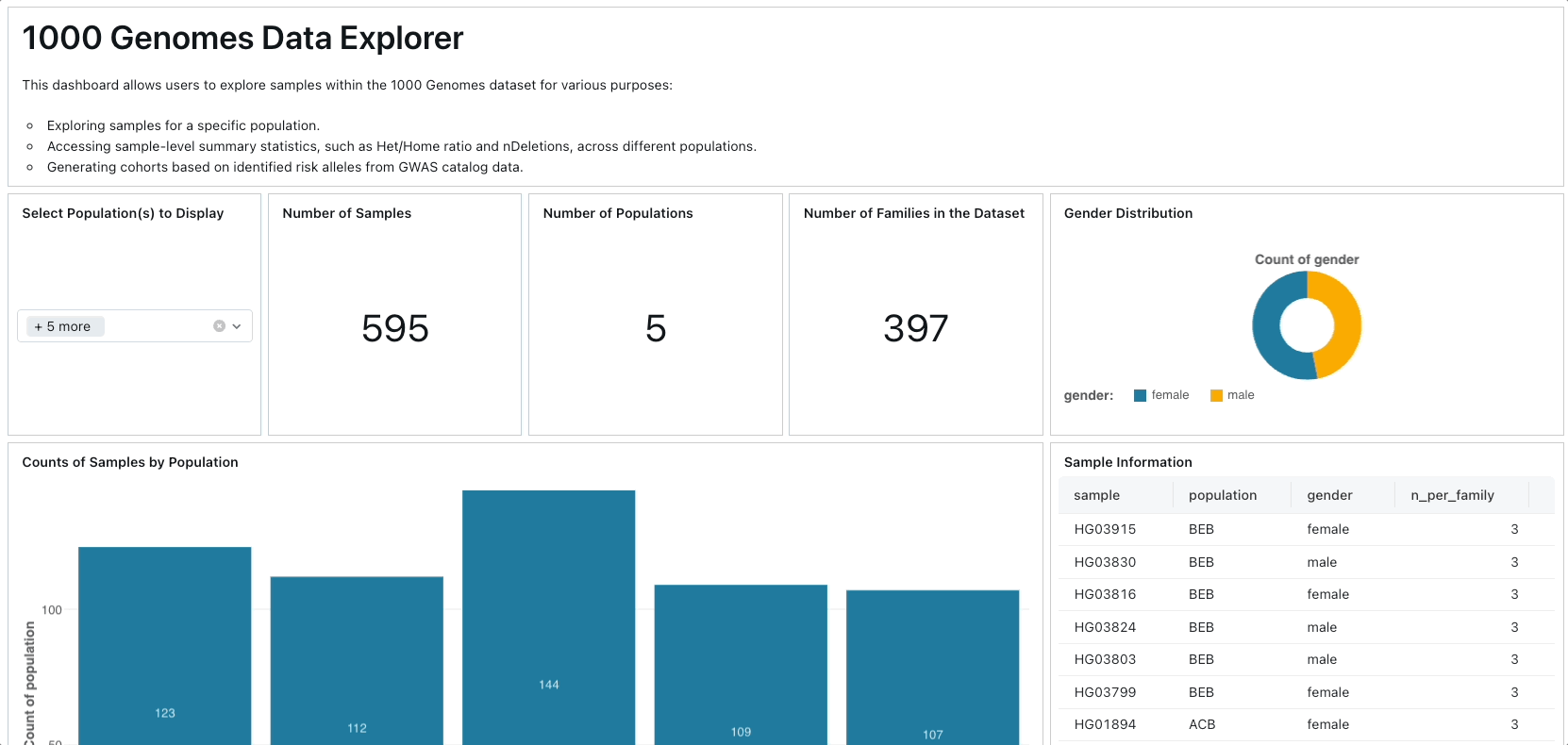
Primarily, we leverage Project Glow to access and ingest 1000 Genomes Project data from public cloud storage. We compute various sample-level and variant-level summary statistics, constructing a database of human genetic variation alongside GWAS catalog data. Subsequently, we develop an interactive dashboard facilitating exploration of genetic variation across different populations and facilitating the identification of samples associated with specific risk alleles for particular traits or diseases.
To create the dashboard, you can simply import the lakeview dashboard's JSON file located in ./resources/1000 Genome Samples Dashboard.lvdash.json in Lakeview.
The 1000 Genomes Project began in 2008 with the aim of mapping human genetic variation. It entailed sequencing the genomes of more than 2,500 individuals from 26 diverse populations worldwide. The project sought to construct a detailed map of genetic distinctions within human DNA. By analyzing genomes from a broad and varied sample, it identified millions of genetic variants, including single nucleotide polymorphisms (SNPs) and structural variations like insertions, deletions, and copy number variations.
Data from the 1000 Genomes Project serves as a vital tool for researchers investigating human genetics. It has been pivotal in numerous studies exploring the genetic underpinnings of complex diseases, contributing to our understanding of human evolution and population history.
The GWAS Catalog, a collaboration between NHGRI and EMBL-EBI, is a curated repository of published genome-wide association studies (GWAS). It houses information on genetic variants, traits, and p-values from these studies, serving as a community resource for data access and analysis tools. The Catalog adheres to FAIR principles, encouraging study identifier citation and offering APIs for data access. Future plans include incorporating unpublished GWAS data. Overall, it is an invaluable asset for researchers exploring the genetic underpinnings of human traits and diseases.
graph LR
subgraph SG1 [EBI]
B[TSV: https://ftp.1000genomes.ebi.ac.uk/vol1/ftp/\ntechnical/working/20130606_sample_info/\n20130606_sample_info.txt]
C[TSV: www.ebi.ac.uk/gwas/api/search/downloads/full]
end
subgraph SG2 [s3://1000genomes/phase1/]
A[VCF: chr22.SHAPEIT2_integrated_phase1_v3]
end
subgraph SG3 [Databricks]
subgraph SG4 [Unity Catalog]
A -- 🧬 Glow \nVCF parser --> AA[glow_chr22_vars]
B --> BB[sample_information]
C --> CC[gwas_catalog_full]
AA --🧬 Glow --> DD[glow_chr22_sample_qc]
AA --🧬 Glow --> EE[glow_chr22_vars]
end
subgraph SG5 [Lakeview]
AA -.-> D(Variant Explorer \n Dashboard)
BB -.-> D
CC -.-> D
DD -.-> D
EE -.-> D
end
end
style SG1 fill:#DCE0E2
style SG2 fill:#DCE0E2
style SG3 fill:#98102A,color:#F9F7F4
style SG4 fill:#EEEDE9
style SG5 fill:#EEEDE9
Please note the code in this project is provided for your exploration only, and are not formally supported by Databricks with Service Level Agreements (SLAs). They are provided AS-IS and we do not make any guarantees of any kind. Please do not submit a support ticket relating to any issues arising from the use of these projects. The source in this project is provided subject to the Databricks License. All included or referenced third party libraries are subject to the licenses set forth below.
Any issues discovered through the use of this project should be filed as GitHub Issues on the Repo. They will be reviewed as time permits, but there are no formal SLAs for support.
© 2024 Databricks, Inc. All rights reserved. The source in this notebook is provided subject to the Databricks License [https://databricks.com/db-license-source]. All included or referenced third party libraries are subject to the licenses set forth below.
| library | description | license | source |
|---|---|---|---|
| Project Glow | an open-source toolkit to enable bioinformatics at biobank-scale and beyond. | Apache 2.0 | https://github.com/projectglow/glow |